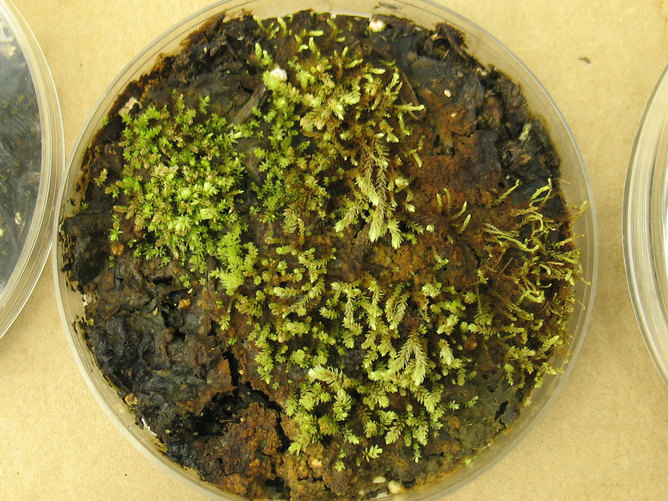
Retreating glaciers are proving to be good news for plant scientists. Underneath one such glacier on Ellesmere Island in Canada, researchers have found plants they believe have regrown after being entombed in the glacier for more than 400 years, since a cold period called the Little Ice Age.
These plants are called bryophyte, a group that includes mosses. They are non-vascular, which means they do not have tissue that distributes resources throughout the plant and they do not reproduce through flowers and seeds. They use spores instead. But they also possess the ability to regrow from tiny fragments of themselves through a process called clonal growth. “This ability makes bryophytes pretty tough,” Andrew Fleming, a plant scientist who was not involved in the study, said.
The discovery reported in the journal Proceedings of the National Academy of Scienes was made by a team led by Catherine La Farge, an expert on bryophytes at the University of Alberta. Because the bryophytes found were not much different from similar variety found in the wild today, La Farge used radio carbon dating to confirm the age of their find.
The plants were trapped during a period known as the Little Ice Age, between the 16th and 19th centuries, when glaciers were growing in size. Arctic glaciers have recently been retreating and, since 2004, the rate of ice melt has increased dramatically. La Farge is hopeful that, in addition to these plants, the melting glaciers will release other interesting flora and fauna of that time.

This discovery does not displace the record of the oldest frozen plant to be regenerated. That belongs to a 32,000 year old specimen of Silene stenophylla, which was regrown by using tissue extracted from its frozen seeds.
These bryophytes are also not the hardiest plants we know. That title belongs to what are commonly known as resurrection plants, which are able to survive extreme dehydration. Some of these are commonly found in deserts, such as Selaginella lepidophylla found in Chihuahuan Desert on the border of Mexico and the US.![]()
First published on The Conversation.
Image credit: Catherine La Farge